Last edit: 05-08-29 Graham Wideman
|
Last edit: 05-08-29 Graham Wideman |
Analyze Format |
| Orientation and Voxel-Order Terminology: RAS,
LAS, LPI, RPI, XYZ and All That Article created: 2003-06-22 |
In dealing with MRI data it's necessary to be familiar with the conventions and terminology used to describe orientation. It's also necessary to be aware of a number of common differences of custom, and various traps for the unwary.
The following is my effort to get all of this straight. It is a work in progress. Even where something's correct in my mind, it's incredibly easy to write it down backwards by mistake. Feel free to send rebuttals to raslasxyz@wideman-one.com
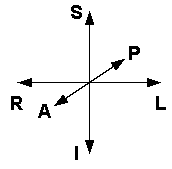
If you're new to the field the first thing you'll need to know is some basic vocabulary regarding directions.
 |
 |
|
| Right | Left | Head orientation reference |
| Anterior | Posterior | |
| Inferior | Superior | |
Notes:
To talk about locations in space in the neighborhood of the brain, we need to be able to talk more precisely about sets of axes, including which direction is positive, and which order we are going to list them when describing a point's coordinates. There are three axes, which could be used in any order, and where either direction could be positive, so that gives us a total of 3 * 2 * 1 * 2 ^ 3 = 48 possible axis schemes.
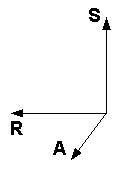
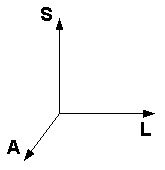
In MRI practice it seems that two of these schemes are most popular:
 |
 |
 |
| "Neurological" convention for axes, aka "RAS" | "Radiological" convention for axes, aka "LAS" | Head orientation reference |
Subtle points:
In some contexts, brain data is placed in an X, Y, Z coordinate system, perhaps like this:
 |
|||
| 1. Would be described as LSA: +X = L +Y = S +Z = A |
2. Would be described as RAS: +X = R +Y = A +Z = S |
3. Would be described as LAS: +X = L +Y = A +Z = S |
Head orientation reference |
Notes:
Why bother renaming perfectly good axes using X, Y, Z, why not just call them R, A, S or whatever? A couple of possible reasons:
A number of MRI file formats involve storing voxel intensities as simply a stream of intensity numbers in to a file in some agreed-upon manner. (Examples: Analyze; AFNI). These formats require recording a number of characteristics of the image file, including voxel order, type of number used for intensity, and other attributes relating to conditions of image acquisition, processing steps that have been performed and so on.
In this discussion we are interested in the topic of voxel order.
In general terms, voxels are stored in sequence traveling along a row, one row after row another, one slice after slice another. But which directions are the rows and slices?
Like the case of the three axes, we again have a choice of 48 possible different storage orders. Here are some:
| Storage order in file | Slice orientation (ambiguous) |
3-Letter "from" Name |
Increasing position in file is in direction of spatial axes... |
| R-L within P-A within I-S | "Axial" | RPI | LAS |
| L-R within P-A within I-S | "Axial" | LPI | RAS |
| R-L within S-I within P-A | "Coronal" | RSP | LIA |
Notes:
"Radiological" and "Neurological" identify two different conventions for the orientation of views of MRI slices. So far as I can tell, they mean the following:
| Convention | Glib descriptions seen everywhere | Slice orientation | Viewer software displays [Note 1] | ||
| Looking towards pt | Patient direction | Screen direction | |||
| Radiological | "Images are viewed as though looking upward
from the feet of the subject." "Right is Left" |
Axial | Superior | Right | Left |
| Anterior | Up | ||||
| Coronal | Posterior | Right | Left | ||
| Superior | Up | ||||
| Sagittal
|
Left? | Anterior | Right? | ||
| Superior | Up | ||||
| Neurological | "Images are viewed as though looking from the
top of the head downward." "Right is right" (Think brain-surgeon-view.) |
Axial
|
Inferior | Right | Right |
| Anterior | Up | ||||
| Coronal | Anterior | Right | Right | ||
| Superior | Up | ||||
| Sagittal
|
Left? | Anterior | Right? | ||
| Superior | Up | ||||
Notes
Given the various schemes for storing data, and the possibility for voxel order to go awry, it's essential to be able to cross-check the actual stored voxel order unambiguously. Surely you can use one of many viewers to do that?
Actually, most viewers go to some effort to read a volume's header file, and orient the view in the most desirable fashion for viewing. This leaves you in the dark regarding the actual order of the data in the file, since it's usually not clear what helpful rotation the viewer has applied.
Here is an example Analyze volume as shown in Chris Rorden's popular MRIcro MRI Viewer. (The data is a test volume available from the FSL folks, that is marked for left and right.)
MRIcro has done a nice job of orienting the views in desirable way. We see that patient right is on the left of the image, so today we're Radiologists. (MRIcro has options for flipping.) But what does this tell us about the actual voxel order on disk?
Here's a screenshot of my format-oblivious Orientation Reality Check viewer that shows the raw data order:
We can see that voxels are ordered (R-L, P-A, I-S) (ie: RPI) in the file. We wouldn't want to study the MRI this way, but it does tell us that what we see in any particular smarter viewer is apt to obscure the raw data order.
Returning to the topic of MRI files and arrays of voxels, some people use letters X, Y and Z when talking about voxel order. Here's a description from SPM regarding the voxel order of files:
[...]after spatial normalisation, the images must be in the following orientation:
- X increases from Left to Right
Y increases from Posterior to Anterior
Z increases from Inferior to SuperiorThis is a right handed coordinate system,
First, let's realize the "images must be in orientation" translates to "voxels must be in this order". Now, from preceding discussion, we can confidently expect that most people will refer to this voxel ordering as LPI, with some others referring to it as RAS. But it's also worth understanding what is the role of X, Y, Z in this picture. Here's my current opinion:
Previous topic notwithstanding, if you were to actually read an MRI file into memory in one big blob and treat it as an array, it would probably be wrong to access it as: Voxels[X, Y, Z]. (Sticking with X being L-R and so on).
That's because in popular computer languages the first index is the slowest-incrementing index into memory. So instead we'd need to access that blob of memory as Voxels[Z, Y, X] (in C/C++: Voxels[Z][Y][X] )
The labels "Radiological" and "Neurological" are sometimes applied to particular orderings of voxels in a file. This is quite a surprise to a database analyst, for whom the physical arrangement of bytes in a file has no more to do with what those bytes mean than the issue of whether they are stored clockwise or counter-clockwise on a hard-drive -- so long as there's metadata on hand to inform us what that order is.
Nonetheless, there has apparently been a history of viewers which slavishly copied bytes out of files and plotted them "verbatim" on screen from left to right, top to bottom, as this is the fastest way to get data from file to screen.
If voxel data is stored in a file from patient-right to patient-left, anterior to posterior, for example) , then when this is "copied" to the screen it places the patient's right on the left of the screen, and anterior at the top, the Radiological preference.
For the Neurological folks who want patient right on the right of the screen, one suspects that there was overwhelming temptation to achieve this effect in early slavishly-Radiological viewers by re-ordering the data. (Or maybe the rationale was to be able to load data into matrices in a particular order -- this would also motivate a desire for right-hand-organized data.)
As a consequence, we have the following terminology used by some:
| Storage order in file | Slice orientation |
3-Letter "from" Name |
Known as |
| R-L within P-A within I-S | "Axial" | RPI | Radiological |
| L-R within P-A within I-S | "Axial" | LPI | Neurological |
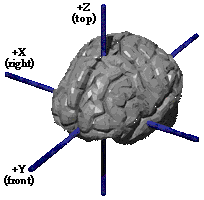
| I took a quick look at how coordinates in the Talairach
atlas are commonly expressed. The
Talairach
Daemon (University of Texas Health Sciences) provides the reference
shown at the right, which seems representative of other Talairach-mentioning
web sites.
Here we see RAS axes, using the letters X Y Z. |
 |
For sake of completeness: it should be noted that choices of axes and indices are only the coarsest issues when it comes to orientation, voxel order and position in space. In addition: